IDEKER LABORATORY
University of California, San Diego
|
Lab Members' Research Projects Network-based Diagnosis and Personalized Medicine Error Analysis and Modeling of DNA microarrays |
| Galactose Utilization as a Model System |
|
We have adopted an integrated, systematic approach to assess, refine, and extend the galactose-utilization (GAL) pathway in the yeast Saccharomyces cerevisiae. First, the GAL pathway was exposed to a battery of twenty perturbations, in which key genes were deleted from the yeast genome and/or key environmental stimuli were altered. Next, whole-genome DNA microarrays and global proteomics were used to track changes in mRNA and protein.
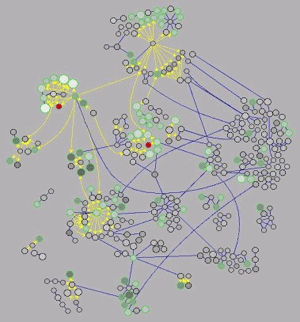
To address expression changes in genes outside of the GAL pathway, we have constructed a network of all known molecular interactions in yeast, drawing data from publicly-accessible databases of protein-protein and protein-DNA interactions. To date, we have been able to explain 20-40% of all gene expression changes, most of which occur in various metabolic pathways, with plausible physical interactions in the network. |